Whether or not bird flu will be transmitted from human to human is still not known. If it does, with no effective vaccine available our only defence will be the antiviral drugs Relenza and Tamiflu.
-
Isolation and crystallisation of neuraminidase on the surface of influenza virus leads to the first effective drugs for controlling flu
-
Chemists identified an active site in the enzyme, which when 'plugged' by a drug stopped the enzyme working and the virus multiplying

Flu, caused by the influenza virus, has plagued mankind for centuries. In 1918-19 the virus, dubbed Spanish flu, killed about 40 million people worldwide. Since then there have been two other major pandemics - in 1957 'Asian flu' killed about two million people and in 1968 'Hong Kong flu' killed about one million people. Of great concern at the moment is a lethal avian influenza virus, or 'bird flu', which has killed millions of domestic poultry, some wild birds and some mammals, including humans. To date, however, people infected with this virus have not been able to infect other people. If this virus does manage to adapt genetically to do this, it will spread like wildfire throughout the world and the antivirals Relenza and Tamiflu look like being our best lines of attack. Here we discover how these drugs were developed and how they can be used effectively.
Viruses take over host cells

Virus particles are essentially nucleic acid (as RNA/DNA) packed into a protein matrix. All viruses have surface antigens (proteins) that can interact with moieties on the surface of their target cells, and are the means by which the virus gets into the cell. The pandemics of 1957 and 1968 were caused by viruses that had surface antigens to which no one had any immunity, allowing the viruses to spread unhindered throughout the world. The destruction of the host cells and the immune response to the virus give rise to the symptoms of the viral infection. In the case of the influenza virus these are high fever, shivering attacks, muscle pains, headache and a dry cough.
In the 1940s George Hirst, working at the Rockefeller Institute, New York, discovered that the influenza virus could attach to receptors on red blood cells in the cold, causing the cells to clump together. When the virus-red cell complexes were warmed to 37°C the cells dispersed and the virus separated. This happened because of a receptor-destroying enzyme on the surface of the virus. This enzyme was eventually identified as a sialidase or neuraminidase by Alfred Gottschalk at the Walter and Eliza Hall Institute, Melbourne, Australia. Specifically, the enzyme cleaved sialic acid from the receptors, destroying their capacity to bind virus particles. Scientists quickly realised that if a 'poison' could be developed that stopped this enzyme working, it might provide a cure for the flu. It took more than 50 years for this idea to be realised.
Progress - haemagglutinin antigen identified
Initially scientists thought that the virus bound to cells via an enzyme-substrate interaction. Slowly, this explanation was shown to be incorrect and it was finally proved by Robin Valentine and myself, working at the National Institute for Medical Research, Mill Hill, London, by using electron microscopy, that there were two distinct antigens with different functions on the surface on the influenza virus. One was the glycoprotein, haemagglutinin, which attaches the virus to receptors on the cells, and the other, the enzyme neuraminidase, destroys these receptors and allows the new virus particles to be released from the cells (see Fig 1).
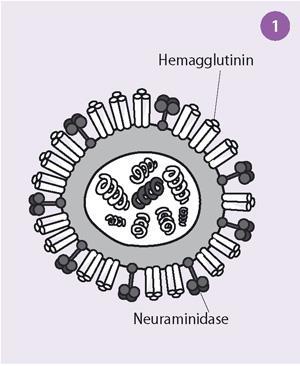
The electron micrographs showed that the influenza virus particles are covered by a layer of surface projections or 'spikes' (see Fig 2). These are of two kinds. One of the spikes, a triangular rod-shaped molecule, has haemagglutinin activity, while the other spike, a mushroom-shaped molecule, with a square box-like head sitting on top of long thin stalk, is the enzyme, neuraminidase. The end of the stalk is hydrophobic which allows the neuraminidase to be attached to the lipid membrane of the virus. Each influenza virus particle is covered by about 2000 haemagglutinin and 500 neuraminidase spikes.
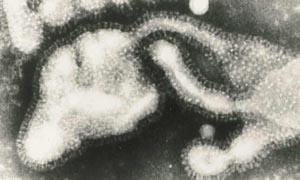
The head of the neuraminidase spike, which contains all the enzymic activity of the molecule, can be released from some influenza virus strains by digesting the virus particles with a protease. The released heads can then be purified and in some cases crystallised. In 1978 I grew the first neuraminidase crystals of influenza virus and, five years later, Peter Colman and his colleagues at the CSIRO Division of Protein Chemistry, Melbourne, solved the three dimensional structure by using x-ray diffraction (Fig 3). This structure revealed that the neuraminidase was a tetramer composed of four identical monomers, each containing a single polypeptide chain. In the centre of each monomer was a deep cleft, or canyon able to bind sialic acid. This was the active, catalytic site of the enzyme.
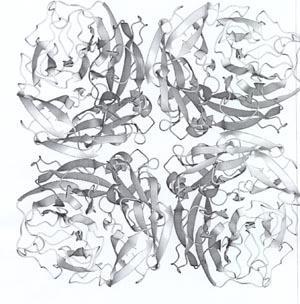
A crucial discovery
It turns out that influenza viruses undergo enormous variation, mainly in the two surface antigens. Because of this influenza vaccines need to be updated every year. To date 16 flu A haemagglutinin subtypes (H1-H16) and nine neuraminidase subtypes (N1-N9) have been found in Nature, some in people, but most in viruses infecting wild water birds. (Note Spanish flu was designated H1N1, Asian flu, H2N2, Hong Kong flu H3N2, and the lethal avian or 'bird flu', H5N1).
Within each subtype there are many different strains. The amino acid sequences of the neuraminidase from a variety of different influenza viruses show enormous variation, some sequences being as much as 75 per cent different. Nevertheless, scattered along the neuraminidase polypeptide at various positions were amino acids that were the same for all strains. When the neuraminidase polypeptide folded into its three dimensional structure, these 'conserved' amino acids were seen to be lining the walls of the catalytic site canyon (Fig 4). This meant that if an inhibitor of the enzyme, a 'plug drug', could be created for one neuraminidase, it would be effective against the neuraminidase of all influenza viruses, even those which had not yet appeared in humans. This was a crucial discovery.
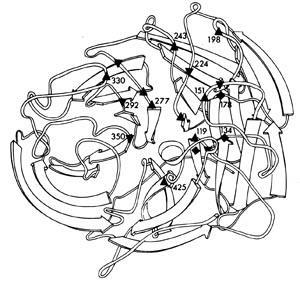
However, there was little point in trying to develop such a 'plug drug' to treat flu infections unless the neuraminidase played an important role in the replication of the virus. Several experiments by Rob Webster and myself (John Curtin School of Medical Research, Canberra), Rudi Rott (Institut für Virologie, Giessen), and Peter Palese and Dick Compass (Mount Sinai School of Medicine, New York), confirmed that it did. Although inhibiting the neuraminidase did not stop the virus from infecting cells, the activity of the enzyme was essential for newly formed virus particles to be released from infected cells and spread in the body to infect other cells. This is because flu viruses infect cells by first binding to receptors on cells via haemagglutinin. The virus then gains entry to the cell by fusing its membrane with the cell membrane, a process also mediated by the haemagglutinin.
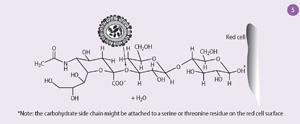
The receptors to which the haemagglutinin binds are composed of chains of sugar molecules with terminal sialic acid residues (Fig 5). The neuraminidase removes the sialic acid from the receptors on cells where the virus is replicating, allowing the newly formed virus particles to escape. But these virus particles also have carbohydrate side chains with terminal sialic acid residues, and these also need to be removed to stop the virus particles sticking together. So the function of the neuraminidase is not only to release the virus from infected cells but also to allow it to spread in the body.
Relenza - the first 'plug drug'
The first neuraminidase inhibitor, or 'plug drug', was created from a knowledge of the crystal structure of a human flu virus neuraminidase, subtype N2. Sialic acid (N-acetyl neuraminic acid, NANA, 1), the substrate of neuraminidase, is itself a mild inhibitor of the enzyme but the dehydrated derivative, deoxy-dehydro-N-acetyl neuraminic acid, DANA (2), the transition state analogue, is a better inhibitor.
Mark von Itzstein and his colleagues at the Victorian College of Pharmacy in Melbourne used computational chemistry techniques to probe the active site of the enzyme in an attempt to design structurally modified derivatives of DANA that would bind tightly to the amino acids in the catalytic site and so prove to be potent and specific inhibitors of the enzyme. They used the software program GRID to determine energetically favourable interactions between various functional groups and residues in the catalytic site canyon. This showed that there was a negatively charged zone in the neuraminidase active site that aligned with the C4 hydroxyl group of DANA. We therefore replaced this hydroxyl with a positively charged amino group; the 4-amino DANA (3) was 100 times better an inhibitor than DANA, owing to the formation of a salt bridge with a conserved glutamic acid (119) in the active site. They then noticed that Glu 119 was at the bottom of a conserved pocket in the active site, which was just big enough to accommodate a more basic functional positively charged group, such as a guanidino group, that was also larger than the amino group.
Von Itzstein's colleague, Wen Yang Wu, synthesised 4-guanidino DANA (4), and found it to be 1000 times better an inhibitor of flu neuraminidase than DANA, and was specific for the flu enzyme, ie it did not inhibit other sialidases found in mammalian cells, some other viruses, bacteria or parasites. 4-Guanidino DANA was named Zanamivir and, after passing clinical trials, is now marketed under the name Relenza.
However, while the presence of the guanidino group made Relenza such a good inhibitor, it also meant it was not orally bioavailable and, if swallowed, was unable to be absorbed from the gut. Relenza therefore had to be administered as a powder that was puffed into the lungs.
Tamiflu tablets
Choung Kim and his colleagues at Gilead Sciences in California then set out to develop a neuraminidase inhibitor that could be swallowed as a pill. Meanwhile, I had organised series of expeditions to Australia's Great Barrier Reef and had isolated, for the first time, a number of avian influenza viruses from wild sea birds, remote from human habitation. One of these viruses had a previously undiscovered neuraminidase, subtype N9, which formed the best crystals of any flu neuraminidase so far examined (Fig 6).

X-ray crystallography of sialic acid bound in this flu neuraminidase showed that the glycerol side chain of the substrate occupied a large pocket in the catalytic site of the neuraminidase protein, made up of a number of hydrophobic amino acids. It also revealed that the C7 position of the sialic acid molecule had no interaction with any of the amino acids in this site. Gilead scientists therefore made carbocyclic sialic acid analogues in which the CHOH group at the C7 position of the glycerol side chain was replaced by an oxygen atom. Then, to create a molecule with hydrophobic groups that would interact well with amino acids in the large hydrophobic pocket, they attached various lipophilic side chains to the oxygen linker that had been introduced at the 7 position.
The carboxylate and acetamido groups corresponding to the same groups on DANA were retained on the new carbocyclic compound and an amino group was introduced at position C4. The final compound chosen, GS4071 (5), had a 3-pentyl side chain and was a potent and specific inhibitor of flu neuraminidase. This compound differed from Relenza in that it was not a sugar, but still suffered from the disadvantage that, because the presence of the amino and carboxyl groups gave it the properties of a zwitterion, it was not orally bioavailable and was unable to be absorbed from the gut.
Gilead scientists solved this problem by converting the carboxyl group to the ethyl ester. The resulting pro-drug, oseltamivir (Tamiflu, 6) can be swallowed as a pill, it is absorbed from the gut and the ester is hydrolysed in the liver to give the active inhibitor that finds its way into the respiratory secretions.
And finally
Relenza and Tamiflu are safe and effective treatments for influenza, but they need to be given early after the first symptoms appear. Six to 12 hours is ideal. In most countries the drugs can only be obtained with a doctor's prescription, and usually the time taken to get a prescription renders them ineffective. Ideally, they should be available 'over-the-counter' in pharmacies without a prescription but only to those people who have genuine influenza. This could be achieved if rapid, sensitive, accurate and inexpensive point-of-care diagnostic tests were also available in pharmacies. Effort is now being put into making this 'test and treat' routine procedure for influenza infections throughout the world.
There is the fear that the influenza virus will develop resistance to the drugs. Inappropriate use will not lead to resistance, but widespread appropriate use might. So far, viruses resistant to Tamiflu have arisen in the human population, but these have not yet spread in the community and the hope is that they never will. Time will tell.
Professor Graeme Laver can be contacted at 3047 Barton Highway, Murrumbateman, NSW 2582, Australia
(e-mail: graeme.laver@bigpond.com)
Further Reading
- G. Laver, Future Virology, 2006, 1, (5), 577.
- E. Garman and G. Laver, Current Drug Targets, 2004, 5, 119.
- A. Moscona, New England J. Med., 2005, 353, 1363.
- M. A. Williams et al, Bioorganic and Med. Chem. Lett., 1997, 7, 1837.






No comments yet